Our Recent Addition to POE Tool Kit
Hybridization of TEDOR and NCX MAS solid-state NMR experiments for simultaneous acquisition of heteronuclear correlation spectra and distance measurements,
T. Gopinath, Songlin Wang, John Lee, Hideki Aihara, and Gianluigi Veglia. Journal of Biomolecular NMR (in press)
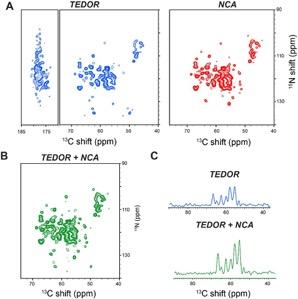
Figure (right): 2D TEDOR and NCA spectra of succinate-acetate permease (SatP) membrane protein acquired simultaneously using the TEDOR-NCA pulse sequence.
Making the Best of NMR Polarization: Polarization optimized experiments (POEs) for high-throughput solid state NMR of proteins
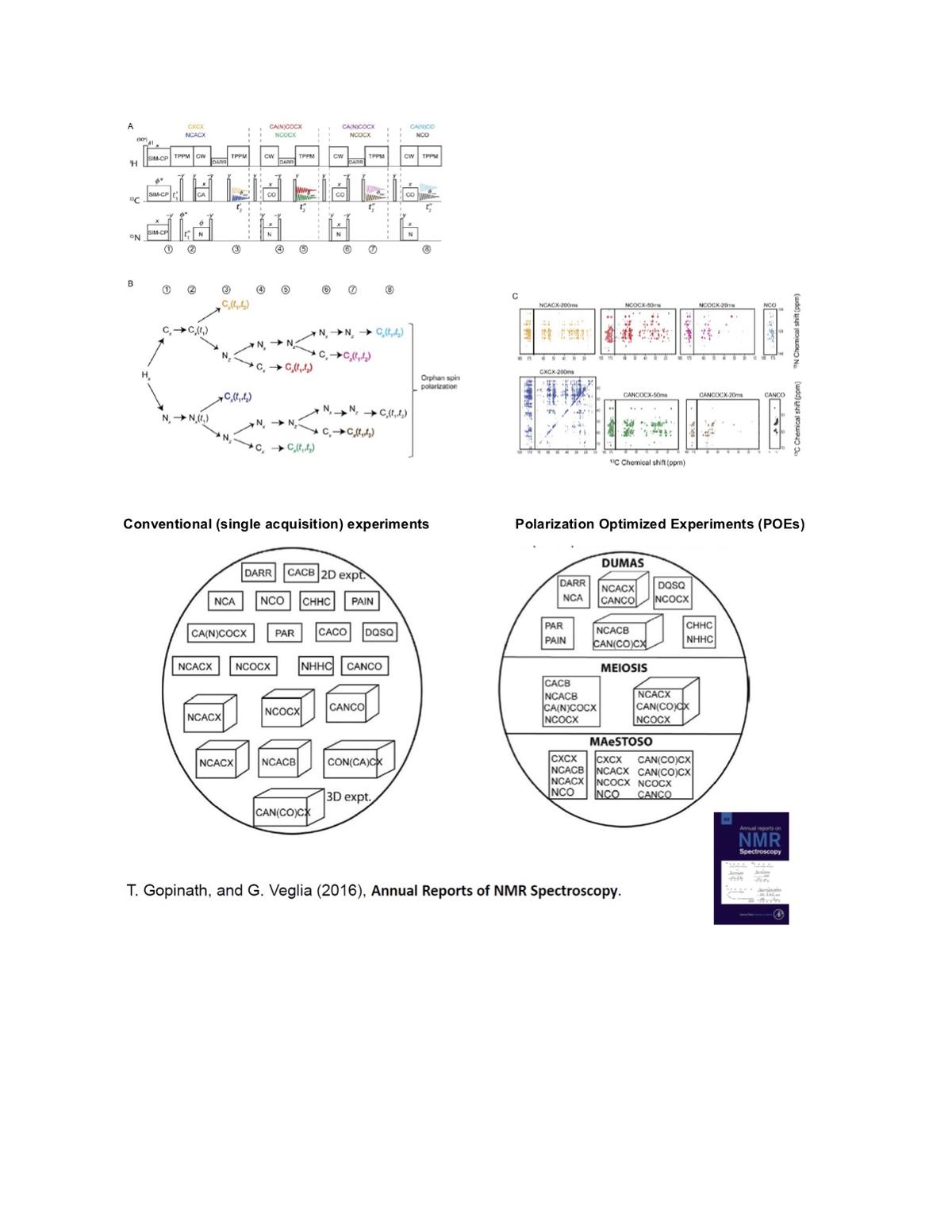
Magic angle spinning (MAS) solid-state NMR (ssNMR) spectroscopy is a major technique for the characterization of the structural dynamics of biopolymers at atomic resolution. However, the low sensitivity of this technique poses significant limitations to its routine application in structural biology. In addition to this, the high lipid-to-protein ratios essential to maintain the functional integrity of membrane proteins also dilute the protein content of MAS samples and substantially increase the experimental time necessary to obtain high-quality spectra. As a result, NMR of membrane proteins requires longer acquisition times compared to fibrils or microcrystalline protein preparations.
To overcome these challenges and reduce experimental time, we developed Polarization Optimized Experiments (POEs), a class of pulse sequences that make the best out of nuclear polarization. POEs enable the acquisition of multiple 2D and 3D NMR spectra simultaneously in a single experiment, resulting in a substantial saving of experimental time. A key element of POEs is the simultaneous cross-polarization (SIM-CP) scheme that matches Hartmann-Hahn conditions for 1 H, 13 C, and 15 N contemporarily to generate an additional 15 N polarization (N z ) that remains stored along the z-axis for several milliseconds due to its relatively long longitudinal relaxation time. Therefore, after the main experiment is recorded (first acquisition), the N z polarization is utilized to generate one or more nD spectra within the same pulse program. The FIDs originating from POEs are usually saved in different memory allocations and processed into separate spectra. The straightforward implementation on commercial NMR spectrometers equipped with only one receiver and probes for bio-solids (Low-E or E-free) made POEs appealing to the broader NMR community. Depending on the types of experiments, POEs use three different strategies (DUMAS, MEIOSIS, and MAESTOSO) to combine various 2D and 3D ssNMR experiments.
Pulse Design Using Evolutionary Algorithm
Interactions of tiny nuclear spin magnets with each other and with the external magnetic fields comprise the whole NMR spectroscopy. Each NMR active nucleus reports the ‘local interactions’ of its local magnetic environment which we can read using the resonance phenomena. Various NMR experiments are devised to read these 'magnetic reports', which essentially depict molecular structure, dynamics, etc. In order to resolve this information, radio-frequency (RF) pulses of specific duration, frequency and phases are required. Often these pulses are associated with experimental inhomogeneities which affect sensitivity and resolution of the experiment. Using genetic algorithm (GA) optimization, we design robust RF pulses for various applications in solution and solid-state NMR. GA optimization is based on natural biological evolution that operates on a population of solutions encoded into a chromosome-like data structure. The evolution of the solutions is obtained by applying recombination operator. As with the natural selection of living organisms, the solutions originating from each generation displaying the greatest fitness have a higher probability to be selected for the next generation. The process continues until the best solution to a specific problem is reached.
